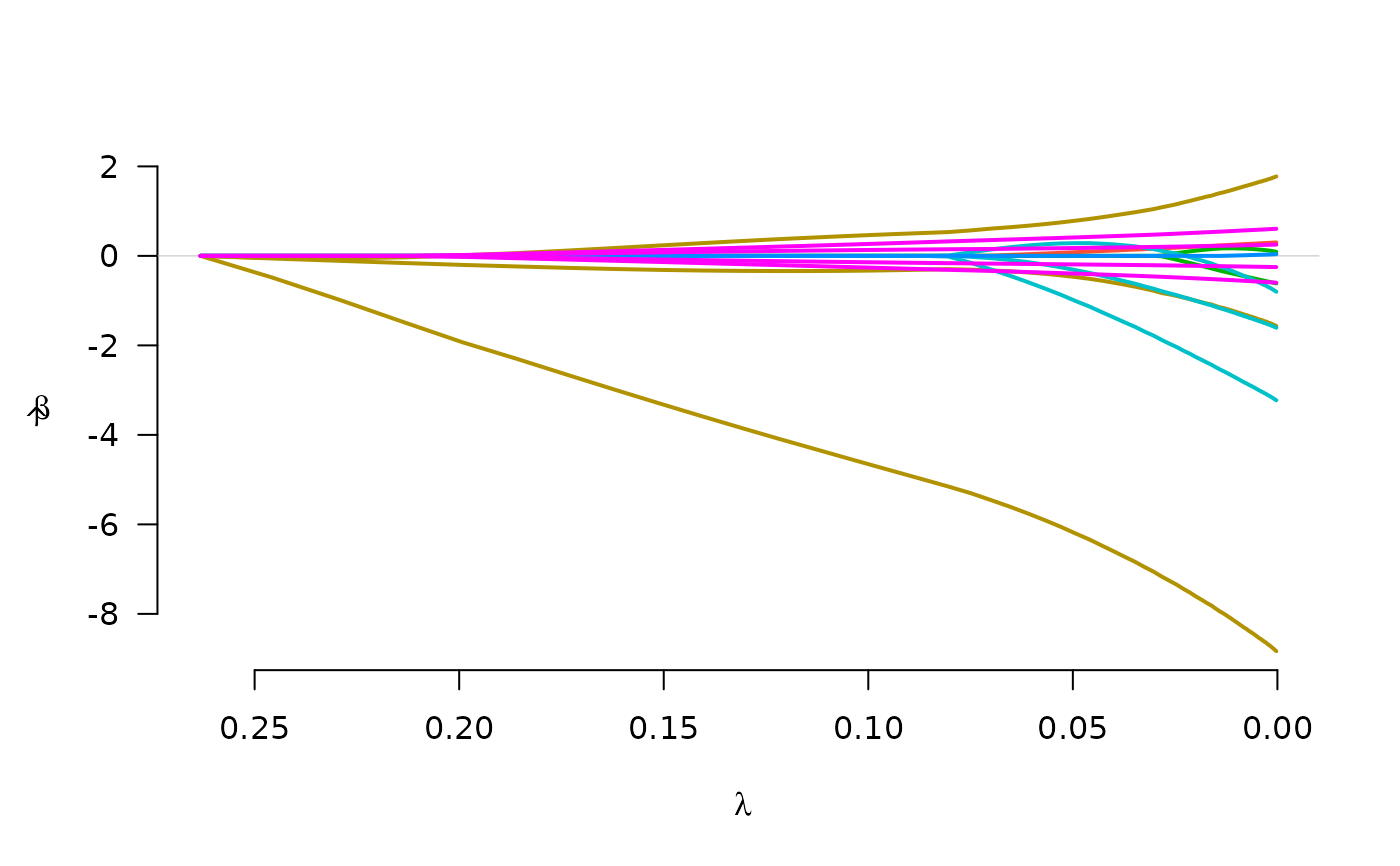
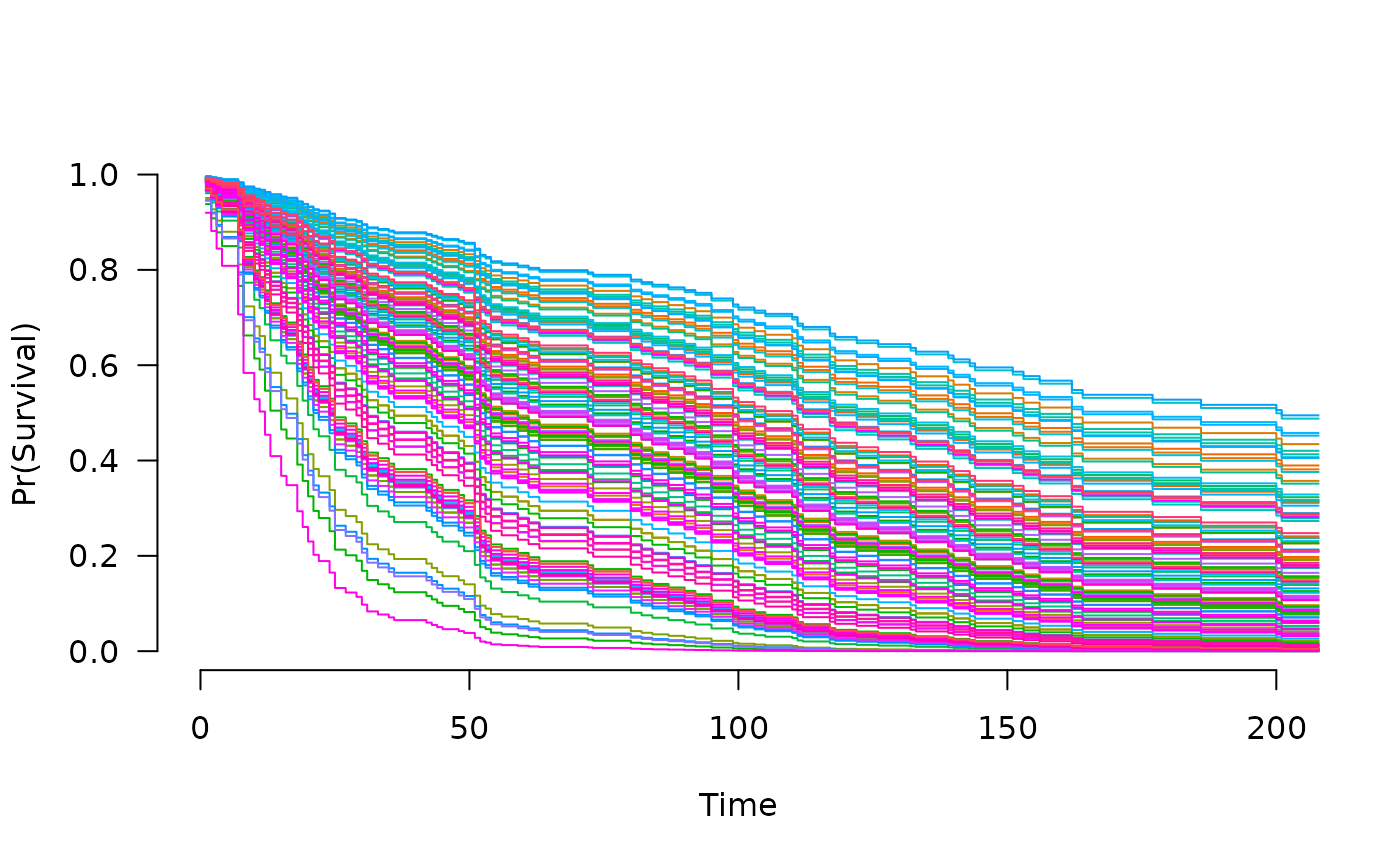
Fit regularization paths for Cox models with grouped penalties over a grid of values for the regularization parameter lambda.
Usage
grpsurv(
X,
y,
group = 1:ncol(X),
penalty = c("grLasso", "grMCP", "grSCAD", "gel", "cMCP"),
gamma = ifelse(penalty == "grSCAD", 4, 3),
alpha = 1,
nlambda = 100,
lambda,
lambda.min = {
if (nrow(X) > ncol(X))
0.001
else 0.05
},
eps = 0.001,
max.iter = 10000,
dfmax = p,
gmax = length(unique(group)),
tau = 1/3,
group.multiplier,
warn = TRUE,
returnX = FALSE,
...
)Arguments
- X
The design matrix.
- y
The time-to-event outcome, as a two-column matrix or
Survobject. The first column should be time on study (follow up time); the second column should be a binary variable with 1 indicating that the event has occurred and 0 indicating (right) censoring.- group
A vector describing the grouping of the coefficients. For greatest efficiency and least ambiguity (see details), it is best if
groupis a factor or vector of consecutive integers, although unordered groups and character vectors are also allowed. If there are coefficients to be included in the model without being penalized, assign them to group 0 (or"0").- penalty
The penalty to be applied to the model. For group selection, one of
grLasso,grMCP, orgrSCAD. For bi-level selection, one ofgelorcMCP. See below for details.- gamma
Tuning parameter of the group or composite MCP/SCAD penalty (see details). Default is 3 for MCP and 4 for SCAD.
- alpha
grpsurvallows for both a group penalty and an L2 (ridge) penalty;alphacontrols the proportional weight of the regularization parameters of these two penalties. The group penalties' regularization parameter islambda*alpha, while the regularization parameter of the ridge penalty islambda*(1-alpha). Default is 1: no ridge penalty.- nlambda
The number of lambda values. Default is 100.
- lambda
A user-specified sequence of lambda values. By default, a sequence of values of length
nlambdais computed automatically, equally spaced on the log scale.- lambda.min
The smallest value for lambda, as a fraction of lambda.max. Default is .001 if the number of observations is larger than the number of covariates and .05 otherwise.
- eps
Convergence threshhold. The algorithm iterates until the RMSD for the change in linear predictors for each coefficient is less than
eps. Default is0.001.- max.iter
Maximum number of iterations (total across entire path). Default is 10000.
- dfmax
Limit on the number of parameters allowed to be nonzero. If this limit is exceeded, the algorithm will exit early from the regularization path.
- gmax
Limit on the number of groups allowed to have nonzero elements. If this limit is exceeded, the algorithm will exit early from the regularization path.
- tau
Tuning parameter for the group exponential lasso; defaults to 1/3.
- group.multiplier
A vector of values representing multiplicative factors by which each group's penalty is to be multiplied. Often, this is a function (such as the square root) of the number of predictors in each group. The default is to use the square root of group size for the group selection methods, and a vector of 1's (i.e., no adjustment for group size) for bi-level selection.
- warn
Return warning messages for failures to converge and model saturation? Default is TRUE.
- returnX
Return the standardized design matrix? Default is FALSE.
- ...
Not used.
Value
An object with S3 class "grpsurv" containing:
- beta
The fitted matrix of coefficients. The number of rows is equal to the number of coefficients, and the number of columns is equal to
nlambda.- group
Same as above.
- lambda
The sequence of
lambdavalues in the path.- penalty
Same as above.
- gamma
Same as above.
- alpha
Same as above.
- deviance
The deviance of the fitted model at each value of
lambda.- n
The number of observations.
- df
A vector of length
nlambdacontaining estimates of effective number of model parameters all the points along the regularization path. For details on how this is calculated, see Breheny and Huang (2009).- iter
A vector of length
nlambdacontaining the number of iterations until convergence at each value oflambda.- group.multiplier
A named vector containing the multiplicative constant applied to each group's penalty.
For Cox models, the following objects are also returned (and are necessary
to estimate baseline survival conditional on the estimated regression
coefficients), all of which are ordered by time on study (i.e., the ith row
of W does not correspond to the ith row of X):
- W
Matrix of
exp(beta)values for each subject over alllambdavalues.- time
Times on study.
- fail
Failure event indicator.
Details
The sequence of models indexed by the regularization parameter lambda
is fit using a coordinate descent algorithm. In order to accomplish this,
the second derivative (Hessian) of the Cox partial log-likelihood is
diagonalized (see references for details). The objective function is
defined to be $$Q(\beta|X, y) = \frac{1}{n} L(\beta|X, y) + $$$$
P_\lambda(\beta)$$
where the loss function L is the negative partial log-likelihood (half the
deviance) from the Cox regression model.
See here for more details.
Presently, ties are not handled by grpsurv in a particularly
sophisticated manner. This will be improved upon in a future release of
grpreg.
References
Breheny P and Huang J. (2009) Penalized methods for bi-level variable selection. Statistics and its interface, 2: 369-380. doi:10.4310/sii.2009.v2.n3.a10
Huang J, Breheny P, and Ma S. (2012). A selective review of group selection in high dimensional models. Statistical Science, 27: 481-499. doi:10.1214/12-sts392
Breheny P and Huang J. (2015) Group descent algorithms for nonconvex penalized linear and logistic regression models with grouped predictors. Statistics and Computing, 25: 173-187. doi:10.1007/s11222-013-9424-2
Breheny P. (2015) The group exponential lasso for bi-level variable selection. Biometrics, 71: 731-740. doi:10.1111/biom.12300
Simon N, Friedman JH, Hastie T, and Tibshirani R. (2011) Regularization Paths for Cox's Proportional Hazards Model via Coordinate Descent. Journal of Statistical Software, 39: 1-13. doi:10.18637/jss.v039.i05