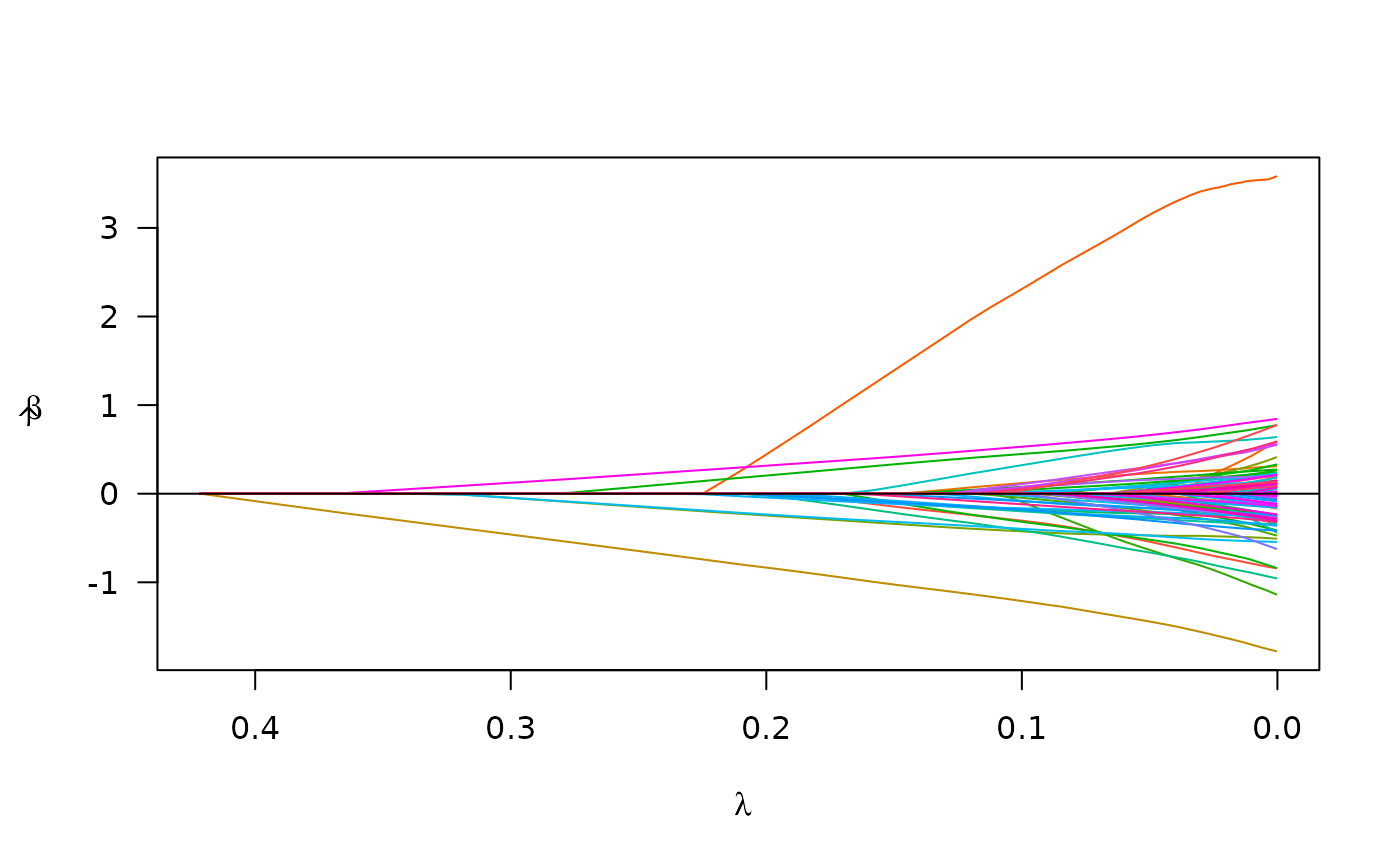
Fit a linear mixed model via penalized maximum likelihood.
Usage
plmm(
design,
y = NULL,
K = NULL,
eta = NULL,
penalty = "lasso",
init = NULL,
gamma,
alpha = 1,
lambda_min,
nlambda = 100,
lambda,
eps = 1e-04,
max_iter = 10000,
dfmax = NULL,
warn = TRUE,
trace = FALSE,
save_rds = NULL,
return_fit = TRUE,
...
)Arguments
- design
The first argument must be one of three things: (1)
plmm_designobject (as created bycreate_design()) (2) a string with the file path to a design object (the file path must end in.rds) (3) amatrixordata.frameobject representing the design matrix of interest.- y
Optional: In the case where
designis amatrixordata.frame, the user must also supply a numeric outcome vector as theyargument. In this case,designandywill be passed internally tocreate_design(X = design, y = y).- K
Similarity matrix used to rotate the data. This should either be: (1) a known matrix that reflects the covariance of y, (2) an estimate (Default is \(\frac{1}{p}(XX^T)\)), or (3) a list with components
sandU, as returned by a previousplmm()model fit on the same data.
Note: If a user provides their ownKmatrix, it is decomposed as provided and will not be scaled. Ifdesignwas created using filebacked data andKis provided by the user, it is possible that a new design matrix containing an intercept will need to be created. This file will be placed in the same directory used to save the final.rdsobject increate_design().- eta
Optional argument to input a specific eta term rather than estimate it from the data. If K is a known covariance matrix that is full rank, this should be 1.
- penalty
The penalty to be applied to the model. Either "lasso" (the default), "SCAD", or "MCP".
- init
Initial values for coefficients. Default is 0 for all columns of X.
- gamma
The tuning parameter of the MCP/SCAD penalty (see details). Default is 3 for MCP and 3.7 for SCAD.
- alpha
Tuning parameter for the Mnet estimator which controls the relative contributions from the MCP/SCAD penalty and the ridge, or L2 penalty.
alpha = 1is equivalent to MCP/SCAD penalty, whilealpha = 0would be equivalent to ridge regression. However,alpha = 0is not supported; alpha may be arbitrarily small, but not exactly 0.- lambda_min
The smallest value for lambda, as a fraction of the maximum lambda. Default is .001 if the number of observations is larger than the number of covariates and .05 otherwise.
- nlambda
Length of the sequence of lambda. Default is 100.
- lambda
A user-specified sequence of lambda values. By default, a sequence of values of length
nlambdais computed, equally spaced on the log scale.- eps
Convergence threshold. The algorithm iterates until the RMSE for the change in linear predictors for each coefficient is less than
eps. Default is1e-4.- max_iter
Maximum number of iterations (total across entire path). Default is 10000.
- dfmax
Maximum number of non-zero coefficients that may enter the model. Default is NULL (no maximum)
- warn
Return warning messages for failures to converge and model saturation? Default is TRUE.
- trace
If set to TRUE, inform the user of progress by announcing the beginning of each step of the modeling process. Default is FALSE.
- save_rds
Optional: if a filepath and name without the
.rdssuffix is specified (e.g.,save_rds = "~/dir/my_results"), then the model results are saved to the provided location (e.g., "~/dir/my_results.rds"). Accompanying the RDS file is a log file for documentation, e.g., "~/dir/my_results.log". Defaults to NULL, which does not save any RDS or log files.- return_fit
Optional: a logical value indicating whether the fitted model should be returned as a
plmmobject in the current (assumed interactive) session. Defaults to TRUE.- ...
Additional optional arguments to
plmm_checks()
Value
A list which includes 18 items:
beta_vals: The matrix of estimated coefficients. Rows are predictors (with the first row being the intercept), and columns are values oflambda.std_Xbeta: A matrix of the linear predictors on the scale of the standardized design matrix. Rows are predictors, columns are values oflambda. Note:std_Xbetawill not include rows for the intercept or for constant features.std_X_details: A list with 9 items:center: The center values used to center the columns of the design matrixscale: The scaling values used to scale the columns of the design matrixns: An integer vector of the nonsingular columns of the original dataunpen: An integer vector of indices of the unpenalized features, if any were specified in the designunpen_colnames: A character vector of the column names of any unpenalized features.X_colnames: A character vector with the column names of all features in the original design matrixX_rownames: A character vector with the row names of all features in the original design matrix; if none were provided, these are named 'row1', 'row2', etc.std_X_colnames: A subset ofX_colnamesrepresenting only nonsingular columns (i.e., the columns indexed byns)std_X_rownames: A subset ofX_rownamesrepresenting rows that passed QC filtering & and are represented in both the genotype and phenotype data sets (this only applies to PLINK data)
std_X: If design matrix is filebacked, the descriptor for the filebacked data is returned usingbigmemory::describe(). If the the data were stored in-memory, nothing is returned (std_Xis NULL).y: The outcome vector used in model fitting.p: The total number of columns in the design matrix (including any singular columns, excluding the intercept).plink_flag: Logical - did the data come from PLINK files?lambda: A numeric vector of the tuning parameter values used in model fitting.eta: A double between 0 and 1 representing the estimated proportion of the variance in the outcome attributable to population/correlation structure.penalty: A character string indicating the penalty with which the model was fit (e.g., 'MCP')gamma: A numeric value indicating the tuning parameter used for the SCAD or MCP penalties. Not relevant for lasso models.alpha: A numeric value indicating the elastic net tuning parameter.loss: A vector with the numeric values of the loss at each value oflambda(calculated on the ~rotated~ scale)penalty_factor: A vector of indicators corresponding to each predictor, where 1 = predictor was penalized.ns_idx: An integer vector with the indices of predictors which were non-singular features (i.e., features which had variation), where feature 1 is the intercept.iter: An integer vector with the number of iterations needed in model fitting for each value oflambdaconverged: A vector of logical values indicating whether the model fitting converged at each value oflambdaK: a list with 2 elements,sandU—s: a vector of the non-zero eigenvalues of the relatedness matrix K (note: K is the kinship matrix for genetic/genomic data; see the article on notation for details)U: a matrix of the eigenvectors of K associated withs
Examples
# using admix data
fit <- plmm(admix$X, admix$y)
s <- summary(fit, idx = 50)
print(s)
#> lasso-penalized regression model with n=197, p=101 at lambda=0.01404
#> -------------------------------------------------
#> The model converged
#> -------------------------------------------------
#> # of non-zero coefficients: 89
#> -------------------------------------------------
plot(fit)