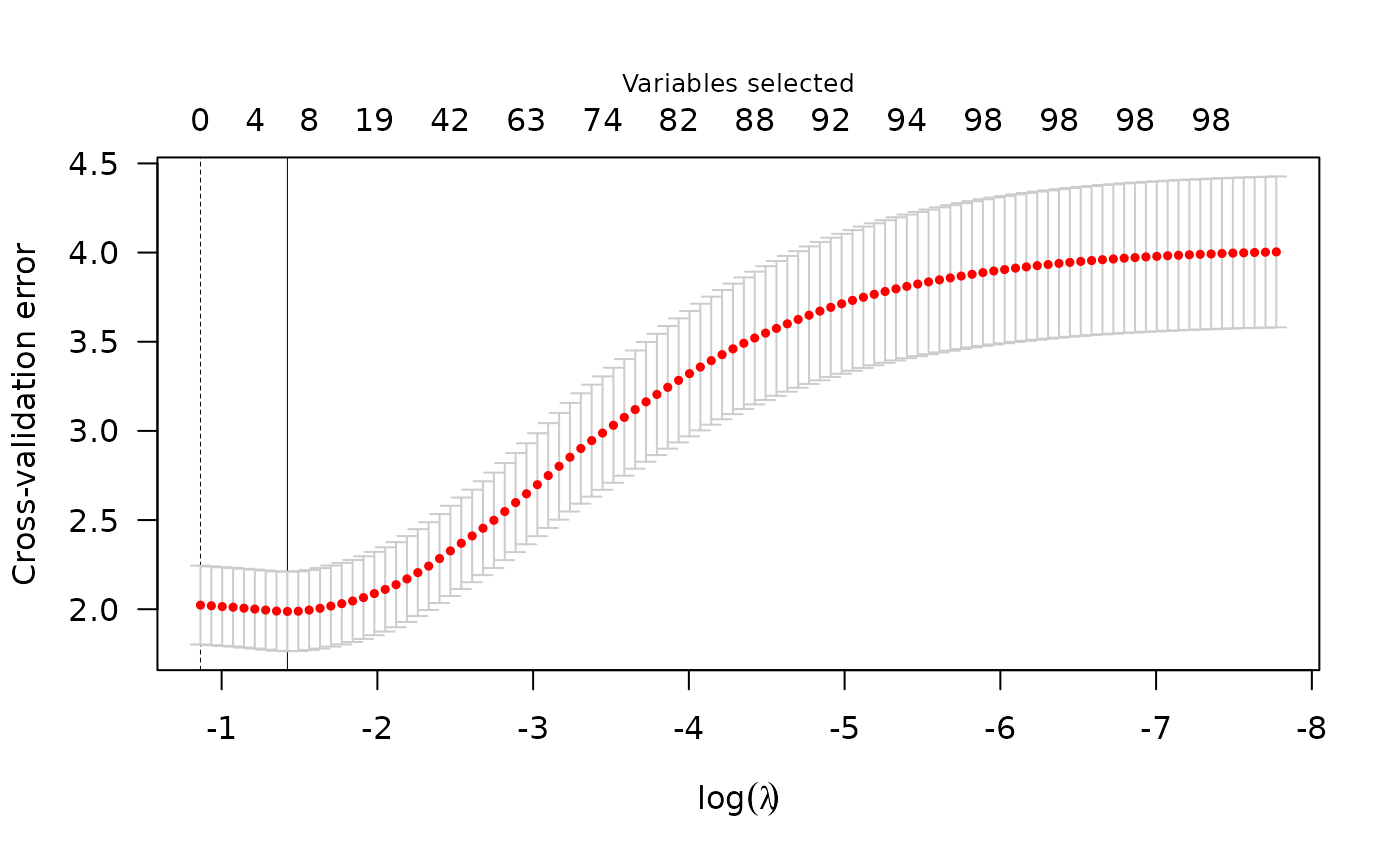
Performs k-fold cross validation for lasso-, MCP-, or SCAD-penalized
linear mixed models over a grid of values for the regularization parameter lambda.
Usage
cv_plmm(
design,
y = NULL,
K = NULL,
eta = NULL,
penalty = "lasso",
type = NULL,
gamma,
alpha = 1,
lambda_min,
nlambda = 100,
lambda,
eps = 1e-04,
max_iter = 10000,
warn = TRUE,
init = NULL,
cluster,
nfolds = 5,
fold = NULL,
seed,
trace = FALSE,
save_rds = NULL,
return_fit = TRUE,
...
)Arguments
- design
The first argument must be one of three things: (1)
plmm_designobject (as created bycreate_design()) (2) a string with the file path to a design object (the file path must end in.rds) (3) amatrixordata.frameobject representing the design matrix of interest- y
Optional: In the case where
designis amatrixordata.frame, the user must also supply a numeric outcome vector as theyargument. In this case,designandywill be passed internally tocreate_design(X = design, y = y).- K
Similarity matrix used to rotate the data. This should either be (1) a known matrix that reflects the covariance of y, (2) an estimate (Default is \(\frac{1}{p}(XX^T)\)), or (3) a list with components
sandU, as returned by a previousplmm()model fit on the same data. Note: if a user provides their own matrix, it is decomposed as provided and will not be scaled.- eta
Optional argument to input a specific eta term rather than estimate it from the data. If K is a known covariance matrix that is full rank, this should be 1. Note: Setting
eta = 1will change the default oftypeto 'lp', as K is always calculated empirically in each fold. This can be overridden by specifyingtype = 'blup', but should be done with caution.- penalty
The penalty to be applied to the model. Either "lasso" (the default), "SCAD", or "MCP".
- type
A character argument indicating what should be returned from
predict.plmm(). Iftype = 'lp', predictions are based on the linear predictor, X beta. Iftype = 'blup', predictions are based on the sum of the linear predictor and the estimated random effect (BLUP). Defaults to 'blup', as this has shown to be a superior prediction method in many applications.- gamma
The tuning parameter of the MCP/SCAD penalty (see details). Default is 3 for MCP and 3.7 for SCAD.
- alpha
Tuning parameter for the Mnet estimator which controls the relative contributions from the MCP/SCAD penalty and the ridge, or L2 penalty.
alpha = 1is equivalent to MCP/SCAD penalty, whilealpha = 0would be equivalent to ridge regression. However,alpha = 0is not supported; alpha may be arbitrarily small, but not exactly 0.- lambda_min
The smallest value for lambda, as a fraction of lambda.max. Default is .001 if the number of observations is larger than the number of covariates and .05 otherwise.
- nlambda
Length of the sequence of lambda. Default is 100.
- lambda
A user-specified sequence of lambda values. By default, a sequence of values of length
nlambdais computed, equally spaced on the log scale.- eps
Convergence threshold. The algorithm iterates until the RMSE for the change in linear predictors for each coefficient is less than
eps. Default is1e-4.- max_iter
Maximum number of iterations (total across entire path). Default is 10000.
- warn
Return warning messages for failures to converge and model saturation? Default is TRUE.
- init
Initial values for coefficients. Default is 0 for all columns of X.
- cluster
Option for in-memory data only:
cv_plmm()can be run in parallel across a cluster using the parallel package. The cluster must be set up in advance usingparallel::makeCluster(). The cluster must then be passed tocv_plmm(). Note: this option is not yet implemented for filebacked data.- nfolds
The number of cross-validation folds. Default is 5.
- fold
Which fold each observation belongs to. By default, the observations are randomly assigned.
- seed
You may set the seed of the random number generator in order to obtain reproducible results.
- trace
If set to TRUE, inform the user of progress by announcing the beginning of each CV fold. Default is FALSE.
- save_rds
Optional: if a filepath and name without the
.rdssuffix is specified (e.g.,save_rds = "~/dir/my_results"), then the model results are saved to the provided location (e.g., "~/dir/my_results.rds"). Defaults to NULL, which does not save the result. Note: Along with the model results, two.rdsfiles ('loss' and 'yhat') will be created in the same directory assave_rds. These files contain the loss and predicted outcome values in each fold; both files will be updated during after prediction within each fold.- return_fit
Optional: a logical value indicating whether the fitted model should be returned as a
plmmobject in the current (assumed interactive) session. Defaults to TRUE.- ...
Additional arguments to
plmm_fit
Value
A list that includes 14 items:
type: The type of prediction used ('lp' or 'blup')cve: A numeric vector with the cross validation error (CVE) at each value oflambdacvse: A numeric vector with the estimated standard error associated with each value ofcvefold: A numericnlength vector of integers indicating the fold to which each observation was assignedlambda: A numeric vector oflambdavaluesfit: The overall fit of the object, including all predictors; this is a list as returned byplmm()min: The index corresponding to the value oflambdathat minimizescvelambda_min: Thelambdavalue at whichcveis minimizedmin1se: The index corresponding to the value oflambdawithin 1 standard error of that which minimizescvelambda1se: The largest value of lambda such thatcveis within 1 standard error of the minimumnull.dev: A numeric value representing the deviance for the intercept-only model. If you have supplied your ownlambdasequence, this quantity may not be meaningful.Y: A matrix with the predicted outcome (\(\hat{y}\)) values at each value oflambda. Rows are observations, columns are values oflambda.loss: A matrix with the loss values at each value of lambda. Rows are observations, columns are values oflambda.estimated_Sigma: Iftype = 'blup', an n x n matrix representing the estimated covariance matrix.
Examples
admix_design <- create_design(X = admix$X, y = admix$y)
cv_fit <- cv_plmm(design = admix_design)
print(summary(cv_fit))
#> lasso-penalized model with n=197 and p=101
#> At minimum cross-validation error (lambda=0.4289):
#> -------------------------------------------------
#> Nonzero coefficients: 0
#> Cross-validation error (deviance): 2.43
#> Scale estimate (sigma): 1.559
plot(cv_fit)